-
GEDmatch
review on 24 January 2017
by Ellen Hinkley

At a Glance
Full Review
GEDmatch is a free genealogy website that allows you to upload the digitised genetic data you receive when you take a test with AncestryDNA, 23andMe, or Family Tree DNA. It was primarily designed to provide another layer of genealogical analysis, to help those who are looking for living relatives aka ‘matches’.
As the ‘matching databases’ maintained by AncestryDNA, 23andMe and Family Tree DNA are kept separate, those looking to find living relatives have historically had to take a test with all three companies to look for all possible matches. Fortunately, GEDmatch allows those who’ve taken a test with just one company to upload their data to its platform, and match with individuals who’ve taken a test with either of the other two (providing those individuals have also uploaded their data to GEDmatch).
As I’d recently taken an AncestryDNA test but had been unable to make much progress with my family tree, I was excited to upload my data to GEDmatch and start searching for living relatives amongst 23andMe and Family Tree DNA users.
Product Expectations
Unfortunately the GEDmatch site has virtually no information on it, but from what I’ve read on various forums, these are the main services they offer: ‘One-to-many matches’, a ‘One-to-one comparison’, an ‘Admixture’ analysis (showing the population groups with whom you share your DNA), ‘Predict eye colour’ and ‘People who match one or both of 2 kits’ (allowing you to search for matches using two of the datasets you’ve uploaded).
I’d also read about ‘Tier 1’ tools which cost $10 to access, and these include the ability to create a dataset for family members who have passed away, and a ‘My Evil twin’ feature which, if your parents have uploaded their data to GEDmatch, lets you create a dataset for a virtual person who’s inherited the genetic variants you could have but didn’t!
My research indicated that GEDmatch maintain a number of projects that analyse their users’ DNA according to their geographic origin, and this helps them refine their ‘Admixture calculators’ (these are what calculate the population groups that you share your DNA with). I was intrigued to see if these calculators, created apart from the three main genetic ancestry companies, would corroborate my AncestryDNA results.
Online Registration
Registration with GEDmatch was straightforward, I just had to provide a few basic details and validate my email address. Once I was able to log in, I selected my zipped AncestryDNA data file for upload and was asked a series of questions, including whether I knew my mitochondrial or Y haplogroups (which I didn’t). I was then required to authorise GEDmatch to make my data available for comparison in their public database.
After uploading my file (it took a few minutes) and returning to the main menu, I could see which ‘batch’ my data file would be processed in (I wasn’t able to use any of the tools before this was done). When I returned to the account a few days later, I could see that on the left-hand side of the main menu under ‘Your DNA Resources’, my kit number and name was now showing.
The Results
The main menu in the GEDmatch account listed numerous tools for me to explore which was quite overwhelming at first. As expected, the following services were now available: ‘One-to-many matches’, ‘One-to-one comparison’, ‘Admixture’, ‘Predict eye colour’ and ‘People who match one or both of 2 kits’.
Results Section: One-to-many matches
When I clicked through to the One-to-many matches section I selected my dataset and was given the option to change the confidence threshold for the matches that would be returned. This was a little over my head, especially as I had to select the threshold by ‘Autosomal’ or ‘X’, and neither option was explained. In the end I chose Autosomal as I understand autosomal DNA makes up the majority of the DNA that we inherit. I was also asked to choose the ‘cM’ (another unexplained term) which I left at ‘7’ which was the default - I later learned that the greater the cM, the greater the certainty of the matches. Upon running the search I was shown a list of 781 matches with names and email addresses which was astounding! My top 20 matches are shown below (excluding their names and email addresses):
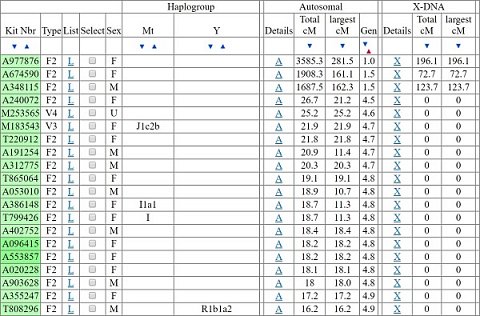
My top 20 matches (excluding their names and email addresses).
I was amazed by the level of detail that was provided with each match, and was surprised that the names and email addresses of every person were provided – most genetic ancestry companies only let you contact your matches using their own messaging system. According to GEDmatch, I shared an autosomal segment that’s at least 7 cM with each match shown, providing it contained at least 700 ‘SNPs’ (aka genetic variants).
Despite understanding just a fraction of what I saw in this section, it was a goldmine for finding matches that I could add to my family tree. And when I later discovered my mitochondrial and Y haplogroups, it was helpful that these had been included against certain matches so I could see if I shared maternal or paternal lineage with them.
I visited this section several times and it was a shame that you couldn’t filter by how recent the matches were, or by the haplogroups they belonged to. Fortunately, the most recent matches are highlighted in green, but you still need to scroll through the list carefully to find them.
On the plus side, it was great to see that virtually everyone I contacted to discuss our shared ancestry replied almost straight away! I expect this is because GEDmatch users are specifically looking for living relatives to connect with.
Results Section: One-to-one comparison
The One-to-one comparison section was primarily for comparing your dataset with the dataset of someone whom you already believe shares your ancestry. I entered a kit number for one of the matches who was quite far down the list, but who appeared to share my maternal lineage (we were in the same mitochondrial haplogroup). Just like for the One-to-many tool I was given several options; I stuck to an Autosomal comparison but this time lowered the cM to 3 as I didn’t expect this match to be very closely related to me. After clicking submit I had to wait a few minutes for the results to be returned (see below).
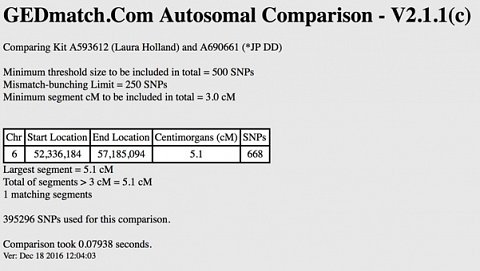

My One-to-one comparison with a match who shares my mitochondrial haplogroup.
As you can see, more detail has been provided about the DNA I share with this match compared to the One-to-many tool, but it’s a shame that so few aspects are explained. I had no idea how I could use the ‘Start Location’ and ‘End Location’ to further my research, and although this information may be valuable if you’re an expert, as someone looking to understand their relationship to key matches, I didn’t find the One-to-one comparison tool to be particularly useful.
Results Section: Admixture
The Admixture section analyses your dataset to identify the population groups that you share your DNA with (the Admixture tool settings are shown below). From what I’d read, as I’m Caucasian British, I needed to select the ‘Eurogenes’ project from the drop-down. As for which process to use, I left the default selected (Admixture Proportions) and continued to the next stage.
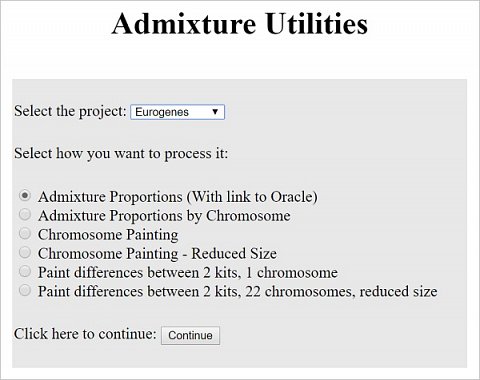
The Admixture tool settings.
I was then asked to confirm the dataset that I wanted to analyse and the calculator that I wanted to use. Unfortunately there were 10 Eurogenes calculators to choose from and I had no way to differentiate between them! I decided to press ahead with the default option (Eurogenes K15) but wished there’d been a way to see how it was different to the others. I was then able to run the calculator and the results were returned as a pie chart (shown below).
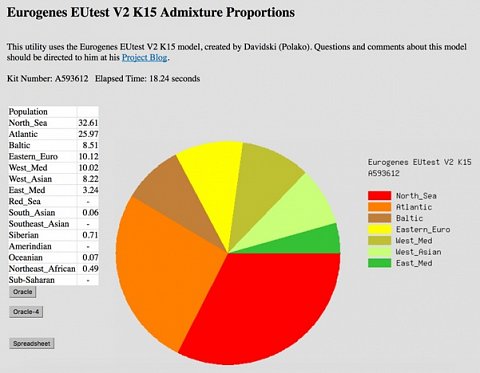
My Admixture results using the Eurogenes K15 calculator.
Upon seeing this chart I got the feeling that the GEDmatch calculators were ‘in development’, and that the results shouldn’t be viewed as definitive. I really liked that GEDmatch returned the name of the administrator for this particular calculator as it felt like I could read his blog, contact him, and really get involved with the Eurogenes project. On the other hand, I found it difficult to derive meaning from the population groups listed as they seemed so vague.
Even though the population groups in this chart were so broad, it was great to see that I shared 10.12% of my DNA with Eastern European populations, and 8.22% with West Asian populations. I felt this corroborated my Romani heritage which has been identified by other tests.
After playing around with this tool I discovered that you can apply additional analyses by using an ‘Oracle’ or ‘Oracle-4’ layer. These layers let me see the population groups I share my DNA with by region (e.g. South Dutch, West German), and I got the feeling that these layers were also in development.
Oracle reported that the top five population groups that had most contributed to my DNA were South Dutch, West German, Southeast English, North German and Southwest English. I’d never seen populations groups reported like this before but I have to say I quite liked the specificity. Oracle went on to report that the ‘mixed’ population I’m most similar to is Southwest English (84.9%) which is spot on! The metric ‘Distance’ was also provided next to each population group, and although I wasn’t sure what this meant, it was clear that the lower the Distance, the higher the percentage of DNA that I shared with that group.
Oracle 4 was similar but reported on my similarity to combinations of two to four population groups. For a mix of two groups I was closest to a 50% Irish/50% West German mixed population, for a mix of three I was closest to 50% Irish/25% Irish/25% Italian, and for a mix of four I was closest to Irish/Irish/Irish/Italian. As you can see, I wasn’t quite able to derive meaning from this analysis, especially as it conflicted with the 84.9% Southwest English result that I’d received earlier.
I found it was important not to take the Admixture results at face value, but to try a number of calculators and look for patterns. If I could ask for two improvements to this section, it would be the addition of a ‘master’ calculator and the ability to filter the results in different ways. Without these, a lot of note taking and spreadsheet work was required to get the most out of this section.
Results Section: People who match one or both of 2 kits
This tool seemed to be designed for helping me work out which of my closest matches can also be matched to my immediate family members. This would depend on those family members uploading a dataset to GEDmatch too, but I could definitely see how this would be useful for working out which branch of the family tree a close match should be added to.
Setting up this tool was similar to setting up the One-to-many tool, except that this time I selected two datasets instead of one: My AncestryDNA dataset and my father’s AncestryDNA dataset. I wasn’t given the option to choose between Autosomal or X, so I presumed matches would be calculated according to autosomal DNA. I was then asked to provide two cM thresholds for the matches; one pertaining to the largest shared segment of DNA, and one pertaining to the total shared segments of DNA. As I wasn’t sure how these would affect the results, I left them both at ‘10’ which were the defaults. Finally, I had to set the “Difference in generations results of 2 kits to common match to disqualify it as a match”. Although there was an explanation of this setting, I couldn’t make any sense of it so I left it at the default ‘99’. After confirming the settings a report was generated which contained three tables: People who match both kits, people who match the 1st kit, people who match the 2nd kit. Each table contained around 300 matches with their email addresses, the top 10 matches who match both kits are listed below (excluding their email addresses):
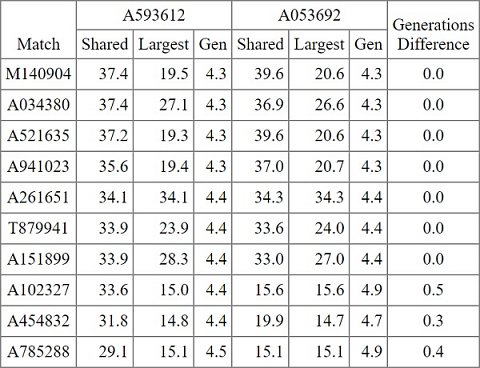
My top 10 matches (excluding their email addresses).
As you can see, the 10 individuals who matched with my father and I all seemed to share ancestors with us four to five generations ago (if I’m interpreting ‘Gen’ correctly). ‘Generations Difference’ appeared to be the difference between ‘Gen’ for each of the two datasets, but I wasn’t really sure why the Generations Difference was meaningful.
I emailed the top 10 (I got several replies straight away) and attempted to add them to the right branch of my family tree. Four generations ago I had 16 ancestors and five generations ago I had 32, so as you can imagine, trying to add the offspring of these individuals to the right branch of my tree takes a lot of correspondence and is quite an undertaking!
One breakthrough I made with this tool was to discover that I shared ancestry with an AncestryDNA match on my father’s side of the family. The individual in question was descended from the inhabitants of Pitcairn Island (a British territory in the Pacific Ocean), and the results confirmed that they shared an ancestor with my father and I, instead of with my mother. Should my immediate family receive additional datasets in the future, I could see how this tool would be extremely useful for adding the matches we find to the right branch of the tree.
Summary
There’s no doubt that GEDmatch is one of the most powerful genealogy tools you can use if you’ve already got a digitised copy of your genetic data from one of the major companies. If you’re dedicated to building your family tree, GEDmatch is a must, especially as the individuals who use it tend to be genealogy enthusiasts who respond to their messages.
A massive downside is that virtually none of the terms used are explained, and unless you have a solid grasp of genetic genealogy, you won’t be able to get the most out of the tools by adjusting their settings.
All in all, GEDmatch is one of the best genealogy tools out there but it’s designed for those with a real enthusiasm for genetic genealogy. If you’re not an expert, be prepared to Google the unexplained terms and search the forums for recommended tool settings – GEDmatch is not for the timid!